Article Search
Volume 27.1
January–April 2024
Full table of contents
ISSN: 1094-8074, web version;
1935-3952, print version
Recent Research Articles
See all articles in 27.1 January-April 2024
See all articles in 26.3 September-December 2023
See all articles in 26.2 May-August 2023
See all articles in 26.1 January-April 2023
Table 1. Summary of specimens used in the training set. F:AMNH refers to the Frick collections of the AMNH; UCMP is the University of California Museum of Paleontology collections. "Hemiauchenia" minima and H. edensis were reassigned by Webb et al. (2008) from Procamelus (Frick, 1921). "H." minima was suggested to be different enough from H. vera that it should have its own generic identification (Webb et al., 1981). Pleiolama vera was reassigned from Hemiauchenia by Webb and Meachen (2004).
|
Taxon |
Location |
Collection |
n |
|
Aepycamelus |
Mixon's Bone Bed, Levy Co., FL |
F:AMNH |
48 |
|
Megatylopus |
Guymon, Texas Co., OK |
F:AMNH |
23 |
|
Procamelus |
Blackhawk Ranch, Contra Costa Co., CA |
UCMP |
6 |
|
Alforjas |
Guymon, Texas Co., OK |
F:AMNH |
24 |
|
"Hemiauchenia" minima |
Mixon's Bone Bed, Levy Co., FL |
F:AMNH |
66 |
|
Hemiauchenia edensis |
Mt. Eden, Riverside Co., CA |
UCMP |
5 |
|
Pleiolama |
Edison Quarry, Sherman Co., KS |
F:AMNH |
9 |
TABLE 2. Eigenvalues, percent of variance explained by each Principal Component, and eigenvectors for the PCA.
TABLE 3. Tukey HSD results for first three PC axes. Taxa not connected by the same letter are significantly different.
|
Taxon |
PC1 |
PC2 |
PC3 |
|||||||
|
Aepycamelus major |
A |
A |
A |
B |
||||||
|
Alforjas |
B |
B |
A |
C |
||||||
|
Megatylopus |
C |
A |
B |
B |
||||||
|
Procamelus |
D |
A |
B |
C |
||||||
|
Pleiolama vera |
E |
A |
A |
B |
||||||
|
"Hemiauchenia" minima |
E |
A |
A |
B |
||||||
|
Hemiauchenia edensis |
E |
A |
B |
A |
B |
C |
||||
TABLE 4. Discriminant function analysis results for training set. Known identity in rows, predicted identity in columns. Correct identifications are in bold, with the number correct followed by the percent correct in parentheses. Incorrect identifications are in blue. The last row represents Mahalanobis distance cutoffs for predictions on unknowns.
TABLE 5. As for Table 4, but reflecting leave-one-out jackknife verification.
TABLE 6. Taxonomic assignments of Thousand Creek unknown camel astragali from each of the three discriminant analyses, accompanied by probability (p) and Mahalanobis Distance. Bold name indicates acceptable identification, while a blue name indicates a rejected identification on the basis of probability (<0.5) or Mahalanobis distance too great. Probabilities and distances are similarly bold or blue to indicate acceptance or rejection. The secondary predictions are listed for each analysis, along with their probabilities and distances; no second prediction is listed if all had p<0.01.
|
Full Dataset |
Genus level |
||||||||||||
|
Loc. No |
Spec. No. |
Prediction |
p |
Dist |
2nd. Pred. |
p |
Dist |
Prediction |
p |
Dist |
2nd |
p |
Dist. |
|
2744 |
70461 |
A. major |
0.95 |
30.6 |
Megatylopus |
0.05 |
36.4 |
Aepycamelus |
0.94 |
30.7 |
Megatylopus |
0.06 |
36.3 |
|
V6570 |
164715 |
Alforjas |
0.63 |
20.3 |
Procamelus |
0.35 |
21.5 |
Alforjas |
0.63 |
20.3 |
Procamelus |
0.34 |
21.6 |
|
2744 |
70463 |
Alforjas |
0.75 |
6.7 |
Procamelus |
0.14 |
10.1 |
Alforjas |
0.77 |
6.6 |
Procamelus |
0.13 |
10.2 |
|
2744 |
164708 |
Alforjas |
0.94 |
12.6 |
Procamelus |
0.05 |
18.4 |
Alforjas |
0.94 |
12.7 |
Procamelus |
0.06 |
18.1 |
|
2744 |
164707 |
Megatylopus |
0.50 |
12.4 |
Procamelus |
0.50 |
12.4 |
Megatylopus |
0.52 |
12.3 |
Procamelus |
0.48 |
12.5 |
|
V78053 |
164712 |
Megatylopus |
0.88 |
27.4 |
Procamelus |
0.08 |
32.3 |
Megatylopus |
0.86 |
27.0 |
Procamelus |
0.11 |
31.2 |
|
V99410 |
158359 |
Megatylopus |
1.00 |
31.6 |
Megatylopus |
1.00 |
31.7 |
||||||
|
V6570 |
31379 |
Procamelus |
0.76 |
19.7 |
Alforjas |
0.23 |
22.0 |
Procamelus |
0.74 |
19.8 |
Megatylopus |
0.01 |
28.9 |
|
2744 |
164709 |
Procamelus |
0.94 |
14.2 |
Megatylopus |
0.06 |
19.7 |
Procamelus |
0.94 |
14.3 |
Megatylopus |
0.06 |
19.7 |
|
1101 |
70346 |
Procamelus |
0.94 |
19.8 |
Megatylopus |
0.05 |
25.5 |
Procamelus |
0.95 |
19.7 |
Megatylopus |
0.05 |
25.6 |
|
2744 |
164728 |
Procamelus |
0.96 |
7.9 |
Alforjas |
0.04 |
14.1 |
Procamelus |
0.95 |
7.9 |
Alforjas |
0.05 |
14.0 |
|
V78050 |
164713 |
Procamelus |
0.98 |
29.6 |
Megatylopus |
0.01 |
38.3 |
Procamelus |
0.98 |
29.7 |
Megatylopus |
0.01 |
38.5 |
|
V99448 |
157285 |
Procamelus |
0.99 |
6.1 |
Procamelus |
0.99 |
6.1 |
||||||
|
2744 |
164725 |
P. vera |
0.32 |
13.3 |
Alforjas |
0.30 |
13.4 |
Pleiolama |
0.36 |
13.4 |
Alforjas |
0.35 |
13.4 |
|
2739 |
164726 |
P. vera |
0.47 |
13.5 |
H. edensis |
0.38 |
13.9 |
Pleiolama |
0.74 |
13.5 |
"Hemiauchenia" |
0.26 |
15.6 |
|
2744 |
70464 |
P. vera |
0.49 |
13.7 |
"H." minima |
0.33 |
14.5 |
Pleiolama |
0.53 |
13.7 |
"Hemiauchenia" |
0.32 |
14.7 |
|
V91097 |
164774 |
P. vera |
0.51 |
3.7 |
Alforjas |
0.23 |
5.3 |
Pleiolama |
0.57 |
3.7 |
Alforjas |
0.26 |
5.3 |
|
V99410 |
158358 |
P. vera |
0.72 |
6.6 |
"H." minima |
0.16 |
9.6 |
Pleiolama |
0.77 |
6.6 |
"Hemiauchenia" |
0.17 |
9.6 |
|
V69107 |
164714 |
"H." minima |
0.45 |
11.6 |
H. edensis |
0.35 |
12.1 |
"Hemiauchenia" |
0.70 |
11.5 |
Pleiolama |
0.30 |
13.2 |
|
V78061 |
164716 |
"H." minima |
0.46 |
4.4 |
H. edensis |
0.44 |
4.6 |
"Hemiauchenia" |
0.83 |
4.3 |
Pleiolama |
0.15 |
7.8 |
|
1100 |
35633 |
"H." minima |
0.48 |
3.2 |
H. edensis |
0.42 |
3.5 |
"Hemiauchenia" |
0.84 |
3.1 |
Pleiolama |
0.15 |
6.5 |
|
V69106 |
164720 |
"H." minima |
0.54 |
4.7 |
H. edensis |
0.43 |
5.2 |
"Hemiauchenia" |
0.96 |
4.6 |
Pleiolama |
0.04 |
11.2 |
|
V6570 |
164727 |
"H." minima |
0.55 |
2.0 |
H. edensis |
0.27 |
3.5 |
"Hemiauchenia" |
0.76 |
2.0 |
Pleiolama |
0.24 |
4.3 |
|
V69106 |
164721 |
"H." minima |
0.55 |
7.6 |
H. edensis |
0.42 |
8.1 |
"Hemiauchenia" |
0.96 |
7.5 |
Pleiolama |
0.04 |
14.0 |
|
V99414 |
158399 |
"H." minima |
0.58 |
2.6 |
H. edensis |
0.34 |
3.7 |
"Hemiauchenia" |
0.88 |
2.5 |
Pleiolama |
0.12 |
6.5 |
|
V99448 |
157286 |
"H." minima |
0.70 |
2.2 |
H. edensis |
0.15 |
5.3 |
"Hemiauchenia" |
0.81 |
2.3 |
Pleiolama |
0.16 |
5.6 |
|
2744 |
164724 |
"H." minima |
0.70 |
3.2 |
H. edensis |
0.21 |
5.6 |
"Hemiauchenia" |
0.88 |
3.2 |
Pleiolama |
0.09 |
7.8 |
|
V69106 |
164723 |
"H." minima |
0.81 |
4.4 |
H. edensis |
0.14 |
7.9 |
"Hemiauchenia" |
0.94 |
4.5 |
Pleiolama |
0.06 |
10.0 |
|
V78075 |
164755 |
H. edensis |
0.37 |
7.2 |
"H." minima |
0.34 |
7.4 |
"Hemiauchenia" |
0.56 |
7.3 |
Pleiolama |
0.44 |
7.7 |
|
V69114 |
164718 |
H. edensis |
0.37 |
15.2 |
"H." minima |
0.16 |
16.9 |
Alforjas |
0.54 |
15.3 |
"Hemiauchenia" |
0.28 |
16.6 |
|
V69106 |
84648 |
H. edensis |
0.51 |
13.8 |
"H." minima |
0.35 |
14.5 |
"Hemiauchenia" |
0.74 |
14.3 |
Pleiolama |
0.26 |
16.4 |
|
V99414 |
157206 |
H. edensis |
0.56 |
9.8 |
"H." minima |
0.30 |
11.1 |
"Hemiauchenia" |
0.71 |
10.8 |
Pleiolama |
0.29 |
12.7 |
|
V99451 |
153817 |
H. edensis |
0.57 |
7.8 |
"H." minima |
0.42 |
8.4 |
"Hemiauchenia" |
0.97 |
8.2 |
Pleiolama |
0.03 |
15.3 |
|
V69106 |
164719 |
H. edensis |
0.72 |
6.6 |
"H." minima |
0.17 |
9.5 |
"Hemiauchenia" |
0.64 |
9.1 |
Pleiolama |
0.36 |
10.3 |
|
2741 |
70417 |
H. edensis |
0.75 |
14.2 |
"H." minima |
0.24 |
16.5 |
"Hemiauchenia" |
0.98 |
16.2 |
Pleiolama |
0.02 |
23.5 |
|
2739 |
70390 |
H. edensis |
0.79 |
16.7 |
"H." minima |
0.19 |
19.6 |
"Hemiauchenia" |
0.91 |
19.2 |
Pleiolama |
0.07 |
24.2 |
TABLE 6 continued.
|
Lumped "Hemiauchenia" |
|||||
|
Prediction |
p |
Dist. |
2nd |
p |
Dist |
|
Aepycamelus |
0.96 |
30.3 |
Megatylopus |
0.04 |
36.5 |
|
Alforjas |
0.69 |
19.8 |
Procamelus |
0.31 |
21.4 |
|
Alforjas |
0.80 |
6.7 |
Procamelus |
0.14 |
10.1 |
|
Alforjas |
0.94 |
12.3 |
Procamelus |
0.05 |
18.1 |
|
Procamelus |
0.50 |
11.7 |
Megatylopus |
0.50 |
11.7 |
|
Megatylopus |
0.75 |
27.0 |
Procamelus |
0.17 |
29.9 |
|
Megatylopus |
1.00 |
31.8 |
|||
|
Procamelus |
0.71 |
19.7 |
Alforjas |
0.28 |
21.6 |
|
Procamelus |
0.96 |
13.1 |
Megatylopus |
0.04 |
19.4 |
|
Procamelus |
0.91 |
19.6 |
Megatylopus |
0.09 |
24.3 |
|
Procamelus |
0.95 |
8.0 |
Alforjas |
0.05 |
14.0 |
|
Procamelus |
0.96 |
29.7 |
Megatylopus |
0.03 |
36.8 |
|
Procamelus |
0.99 |
6.1 |
Alforjas |
0.01 |
16.0 |
|
"Hemiauchenia" |
0.50 |
13.4 |
Alforjas |
0.49 |
13.5 |
|
"Hemiauchenia" |
0.98 |
14.9 |
Alforjas |
0.02 |
23.2 |
|
"Hemiauchenia" |
0.75 |
14.2 |
Alforjas |
0.24 |
16.4 |
|
Alforjas |
0.50 |
5.3 |
"Hemiauchenia" |
0.50 |
5.3 |
|
"Hemiauchenia" |
0.82 |
8.7 |
Alforjas |
0.18 |
11.7 |
|
"Hemiauchenia" |
1.00 |
11.4 |
|||
|
"Hemiauchenia" |
0.98 |
4.4 |
Alforjas |
0.02 |
11.8 |
|
"Hemiauchenia" |
0.99 |
3.2 |
|||
|
"Hemiauchenia" |
1.00 |
4.9 |
|||
|
"Hemiauchenia" |
1.00 |
1.9 |
|||
|
"Hemiauchenia" |
1.00 |
7.8 |
|||
|
"Hemiauchenia" |
0.99 |
2.7 |
|||
|
"Hemiauchenia" |
0.96 |
2.4 |
Alforjas |
0.04 |
8.6 |
|
"Hemiauchenia" |
0.96 |
3.4 |
Alforjas |
0.04 |
10.0 |
|
"Hemiauchenia" |
1.00 |
4.7 |
|||
|
"Hemiauchenia" |
0.99 |
7.0 |
|||
|
Alforjas |
0.64 |
15.3 |
"Hemiauchenia" |
0.36 |
16.5 |
|
"Hemiauchenia" |
1.00 |
14.3 |
|||
|
"Hemiauchenia" |
1.00 |
10.8 |
|||
|
"Hemiauchenia" |
1.00 |
8.5 |
|||
|
"Hemiauchenia" |
1.00 |
8.9 |
|||
|
"Hemiauchenia" |
1.00 |
16.5 |
|||
|
"Hemiauchenia" |
0.97 |
19.5 |
Alforjas |
0.03 |
26.8 |
 Edward Byrd Davis
Edward Byrd Davis
University of Oregon Museum of Natural and Cultural History
and Department of Geological Sciences
1680 East 15th Avenue, Eugene, Oregon 97403 USA
edavis@uoregon.edu
Edward Byrd Davis is the Fossil Collections Manager for the University of Oregon Museum of Natural and Cultural History. He received his B.S. from the University of Tennessee, Knoxville, and his Ph.D. from the University of California, Berkeley. He divides his time between museum curatorial nightmares, teaching in the UO Geology Department, and paleomammalogy research. His primary research interests lie in geographic range response to climate change over shallow and deep time and in ruminant artiodactyl evolution.

 Brianna K. McHorse
Brianna K. McHorse
University of Oregon Clark Honors College
and Department of Biology
1293 University of Oregon
Eugene, Oregon 97403 USA
Current address:
Harvard University Department of Organismic and Evolutionary Biology
26 Oxford Street
Cambridge, Massachusetts 02138 USA
bmchorse@fas.harvard.edu
Brianna K. McHorse graduated from the Robert D. Clark Honors College at the University of Oregon with a degree in Biology. Beginning in fall 2013 she will be a graduate student in the Department of Organismic and Evolutionary Biology at Harvard University, where she will focus on evolutionary functional morphology and mammalian locomotor biomechanics. She has been funded as a Goldwater Scholar investigating the relationship between conformation and competition performance in three-day event horses and as a National Science Foundation Graduate Research Fellow.
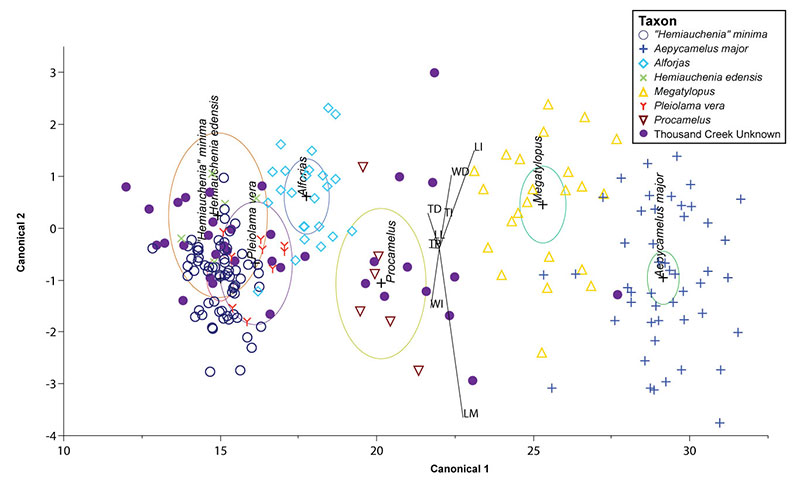
FIGURE 1. Illustration of astragali showing the dimensions used in this study. After DeGusta and Vrba (2003). 1.1, medial view; 1.2, lateral view; 1.3, anterior view. Abbreviations: LM= medial length; LI= intermediate length; LL= lateral length; TD= distal thickness; TI= intermediate thickness; TP= proximal thickness; WD= distal width; WI= intermediate width.
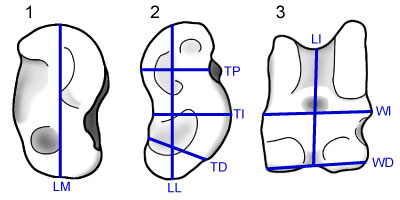
FIGURE 2. Principal Components of known astragali and Thousand Creek astragali illustrating the array of specimens along PC1, but no obvious differences along PC2. While a qualitative examination of PC2 produces no distinction, Tukey's HSD test indicates two separate groups (see Table 3). The Thousand Creek specimens show a single clear break in size, but without the DFA it would be impossible to quantify the certainty of group membership of specimens near the breaks between taxa.
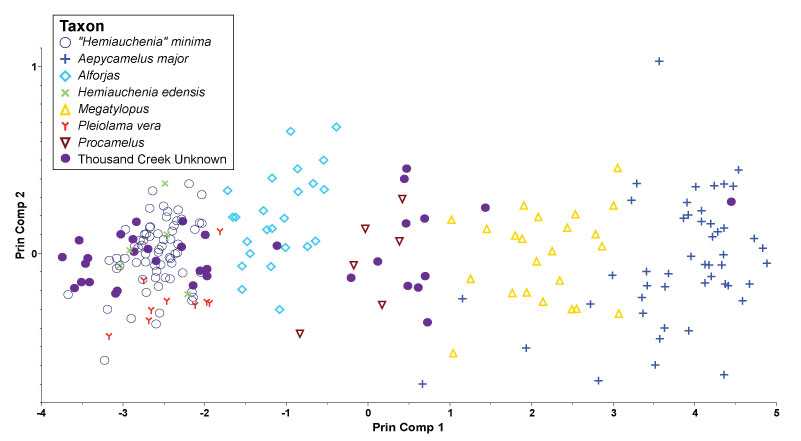
FIGURE 3. First two discriminant axes from species-level analysis. Camel taxa from the training set are indicated by the same shapes and colors as Figure 2. Ellipses show the 95% confidence interval of the true mean.
A method for improved identification of postcrania from mammalian fossil assemblages: multivariate discriminant function analysis of camelid astragali
Edward Byrd Davis and Brianna K. McHorse
Plain Language Abstract
Skulls and teeth are often the most useful bones for identifying mammal fossils because their complex shapes make them easy to distinguish among species. However, many fossil sites consist mostly of limb or foot bones, making it difficult to understand how many different species of mammals lived there. If we can correctly identify a fossil using these well-preserved foot and limb bones, we can use better estimates of mammalian diversity for larger-scale studies that look at ecosystem changes through time. The astragalus, one of the ankle bones, is particularly useful for such work because they preserve extremely well in the fossil record and measurements of certain parts can distinguish between groups of cloven-hooved mammals (artiodactyls). The Thousand Creek fossil sites of Nevada (approximately 8 million years old) represent one such site with abundant foot and limb bones but relatively few skulls and teeth, leaving us unsure about how many different kinds of mammalian species lived there, including camels. We use eight linear measurements on the camel astragali to identify the Thousand Creek camels. First we measured the astragali of previously identified camels from the same time period, forming a data set that established the typical shapes and sizes of those known camels. We used this data set of known camels to perform a discriminant function analysis (DFA), a mathematical way of predicting what group a specimen belongs to on the basis of its measurements. The DFA uses measurements of the known camels to figure out the differences in shapes and sizes among these groups. Next we put our measurements of the Thousand Creek camels into the discriminant function, which determines which taxonomic group or groups the unknown camel probably belongs to. The discriminant function classified successfully four groups of camels: Alforjas, Procamelus, one that is very like Hemiauchenia, and a large Megatylopus-like one. Adding more known camels to our data set will allow more certainty in future identifications, and our discriminant function can also be used to identify camels from other faunas of similar age. Using DFA is more rigorous than simply plotting relationships and drawing circles or lines to separate the groups, especially because it can be used in other studies and refined as more known samples are acquired.
Resumen en Español
Un método para mejorar la identificación de postcráneos en las asociaciones fósiles de mamíferos: análisis de función discriminante de astrágalos de camélidos
Los ejemplares craneodentales, con abundantes caracteres diagnósticos, constituyen normalmente el mejor material para identificar los fósiles de mamíferos a nivel genérico y específico, pero ¿qué se puede hacer con las asociaciones compuestas fundamentalmente por postcráneos desarticulados? En localidades en las que faltan restos diagnósticos, la identificación precisa del material postcraneal puede mejorar las estimaciones de diversidad de mamíferos para estudios a mayor escala. En particular, los astrágalos, que frecuentemente se encuentran bien conservados, han demostrado tener una utilidad diagnóstica en artiodáctilos. La fauna de Thousand Creek de Nevada (~8 Ma) es una de esas asociaciones ricas en material postcraneal pero con una diversidad desconocida para muchos taxones, incluyendo los camélidos. Hemos usado el análisis de función discriminante de ocho medidas lineales en astrágalos de camélidos contemporáneos con afinidad taxonómica conocida para elaborar un conjunto de instrucciones que puede ser utilizado para asignar taxones en el material de camélidos de Thousand Creek. La función discrimanante identifica, como mínimo, cuatro clases de camellos: "Hemiauchenia", Alforjas, Procamelus y ?Megatylopus. La adición de más ejemplares al conjunto de instrucciones puede mejorar la certidumbre y precisión en futuros trabajos, incluyendo la identificación de camélidos en otras faunas de edad similar. Para un mejor tratamiento estadístico y una mayor comodidad en usos futuros recomendamos el empleo del análisis de función discriminante antes que los análisis cualitativos de biplots para separar y diagnosticar taxones.
PALABRAS CLAVE: análisis de función discriminante; Camelidae; Thousand Creek; astrágalo; Mioceno; Hemphilliense
Traducción: Miguel Company
Résumé en Français
Une méthode pour une meilleure identification du postcrânien dans les assemblages fossiles mammaliens : analyse de fonction discriminante multivariée des astragales de camélidés
Les spécimens crânio-dentaires, riches en caractères, sont souvent le meilleur matériel pour identifier les fossiles de mammifères au niveau générique ou spécifique, mais que peut-on faire avec les nombreux assemblages principalement composés de postcrânien dissocié ? Dans une localité manquant de reste typiquement diagnostique, une identification juste du matériel postcrânien peut améliorer les mesures de diversité des mammifères dans les études à grande échelle. Les astragales, en particulier, sont souvent bien préservés et leur utilité diagnostique chez les artiodactyles a été démontrée. La faune de Thousand Creek au Nevada (~8 Ma) représente un de ces assemblages riche en postcrânien mais dont la diversité de ses nombreux taxa reste inconnue, entre autre pour les camélidés. Nous utilisons une analyse de fonction discriminante (DFA) de huit mesures linéaires sur les astragales de camélidés actuels dont les affinités taxinomiques sont connues afin de produire un jeu de référence qui peut être utilisé pour identifier le matériel de camélidés de Thousand Creek. La fonction discriminante identifie, au minimum, quatre catégories de chameaux : "Hemiauchenia", Alforjas, Procamelus, et ?Megatylopus. Un ajout de spécimens, incluant l'identification de camélidés d'autres faunes d'âge similaire, dans le jeu de référence pourrait améliorer la justesse et la précision pour des travaux ultérieurs. Pour un meilleur usage statistique et une utilisation future facilitée, nous recommandons d'utiliser la DFA plutôt que les analyses qualitatives de diagrammes biplots pour séparer et diagnostiquer les taxons.
Mots clés : analyse de fonction discriminante ; Camelidae ; Thousand Creek; astragale; Miocène; Hemphillien
Translator: Olivier Maridet
Deutsche Zusammenfassung
Eine Methode zur besseren Erkennung von Postcranialmaterial aus fossilen Säugetier-Assemblagen: multivariate Diskriminanzfunktionsanalyse von Kamel-Astragali
Merkmalsreiche craniodentale Stücke sind oft das beste Material zur Bestimmung von Säugetierfossilien auf Gattungs-oder Artebene. Doch was ist mit den vielen Assemblagen zu tun, die hauptsächlich aus dissoziiertem Postcranialmaterial bestehen? Bei Lokalitäten ohne typische diagnostische Überreste, kann die genaue Identifizierung des Postcranialmaterials die Maße zur Säugetierdiversität für Untersuchungen auf breiterer Basis verbessern. Besonders Astragali sind oft gut erhalten und haben gezeigt, dass sie diagnostischen Nutzen bei Atriodactylen haben. Die Thousand Creek Fauna von Nevada (~8 Ma) repräsentiert eine solche Fauna, die reich an Postcranialmaterial ist, bei der jedoch die Diversität vieler Taxa – einschließlich der Kamele – unbekannt ist. Mit Diskriminanzfunktionsanalyse (DFA) von acht linearen Maßen bei zeitgleichen Kamel-Astragali mit bekannter taxonomischer Affinität, erstellten wir einen Trainingssatz mit dem Taxa zum Kamelmaterial von Thousand Creek zugeordnet werden können. Die Diskriminanzfunktion identifiziert mindestens vier Kamelklassen: "Hemiauchenia", Alforjas, Procamelus und ?Megatylopus. Weitere dem Trainingssatz zugefügte Stücke würden wahrscheinlich die Sicherheit und die Genauigkeit zukünftiger Arbeiten erhöhen, einschließlich der Identifikation von Kamelen anderer Faunen und mit ähnlichem Alter. Für die beste statistische Praxis und zukünftige Benutzerfreundlichkeit, empfehlen wir eher DFA anstatt qualitative Analyse von Biplots um Taxa zu trennen und zu bestimmen.
Schlüsselwörter: Diskriminanzfunktionsanalyse; Camelidae; Thousand Creek; Astragalus; Miozän; Hemphilium
Translator: Eva Gebauer
Arabic

Translator: Ashraf M.T. Elewa
-
-
-
Review: The Princeton Field Guide to Mesozoic Sea Reptiles
 The Princeton Field Guide to Mesozoic Sea Reptiles
The Princeton Field Guide to Mesozoic Sea ReptilesArticle number: 26.1.1R
April 2023


 Poster Winners 2024
Poster Winners 2024