Article Search
Volume 27.1
January–April 2024
Full table of contents
ISSN: 1094-8074, web version;
1935-3952, print version
Recent Research Articles
See all articles in 27.1 January-April 2024
See all articles in 26.3 September-December 2023
See all articles in 26.2 May-August 2023
See all articles in 26.1 January-April 2023
SUPPLEMENTAL MATERIAL
The Matlab code and files used to generate Figure 2, Figure 5, Figure 6, Figure 7 and Figure 8. To run, download and unzip the Segmentation Demo file folder. In Matlab, navigate to the Segmentation Demo file folder and open the folder. There are three demos named demo1_preprocess, demo2_segment, and demo3_postprocess. You must run the demos in order. To run the demo, simply type the demo’s name in the command prompt and hit enter (see zipped file).
FIGURE 1. Animation of 3-D reconstruction of a crinoid holdfast from CT z-sections. Eucalyptocrinites crassus Waldron, Indiana. Field Museum P569. Click on image to activate.
FIGURE 2. 1. The original image of the crinoid specimen (Eucalyptocrinites crassus, Silurian, Waldron Shale, Waldron, IN. USNM S 366). The black and white scale bar equals 1 cm. 2. Image subset or tile used in the demonstrations throughout this paper. The tile is 512 pixels by 512 pixels. 3. The steps of digital extraction through image segmentation. All photos underwent pre-processing, processing, and post-processing. However, the actual techniques used for each step varied and, in some cases, multiple steps were used. For example, for many of the images both CC analysis (CCA) and binary image morphology were used to improve the quality of the thresholding. A photo of a crinoid root segment is used in here to provide examples of the products of each step.
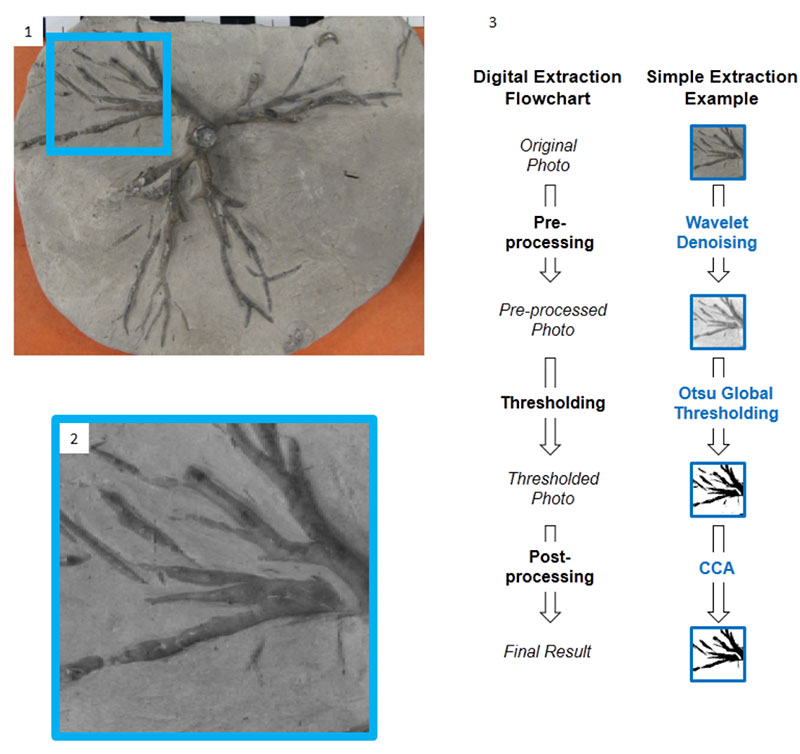
FIGURE 3. Left column: original images of crinoid holdfast. On the right: cleanly segmented images produced from images in the left column by combining several traditional and computer techniques. These segmented images can be used in morphometrics. 1. USNM S366. Frank Springer collection, Waldron, Indiana. 2. Field Museum, U.C. 56306, Waldron, Indiana. 3. USNM 42233, Ulrich collection, Waldron, Indiana. 4. USNM 42233, Ulrich collection, Waldron, Indiana.
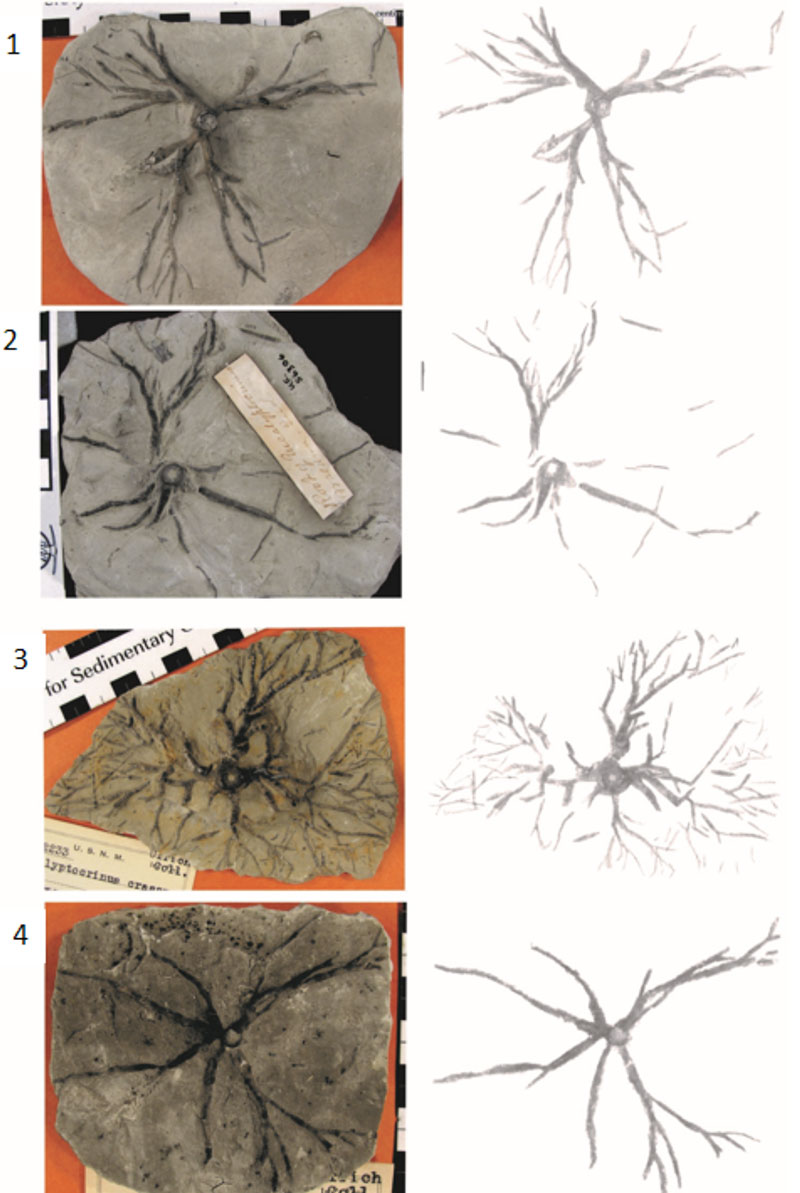
FIGURE 4. 1. Scanned image of a book (American Geosciences Institute Data Sheets, 1982). 2. Grayscale frequency distribution of image shown in 4.1. Because the background and foreground are clearly distinct, Otsu’s method works well even when applied globally with no pre-processing. The red line is the threshold chosen to segment the image. 3. Image in 4.1 treated with Otsu’s method. The details of the image shown in 4.1 are very well preserved in the treated image. 4. Checkerboard shadow illusion developed by Adelson at the Massachusetts Institute of Technology as part of the joint collaboration between the Department of Brain and Cognitive Sciences and the Computer Science and Artificial Intelligence Laboratory (CSAIL). 5. The white squares within the shadow area are incorrectly identified as dark squares by global Otsu’s method without preprocessing (Images from AGI Data Sheets and CSAIL are used with permission).
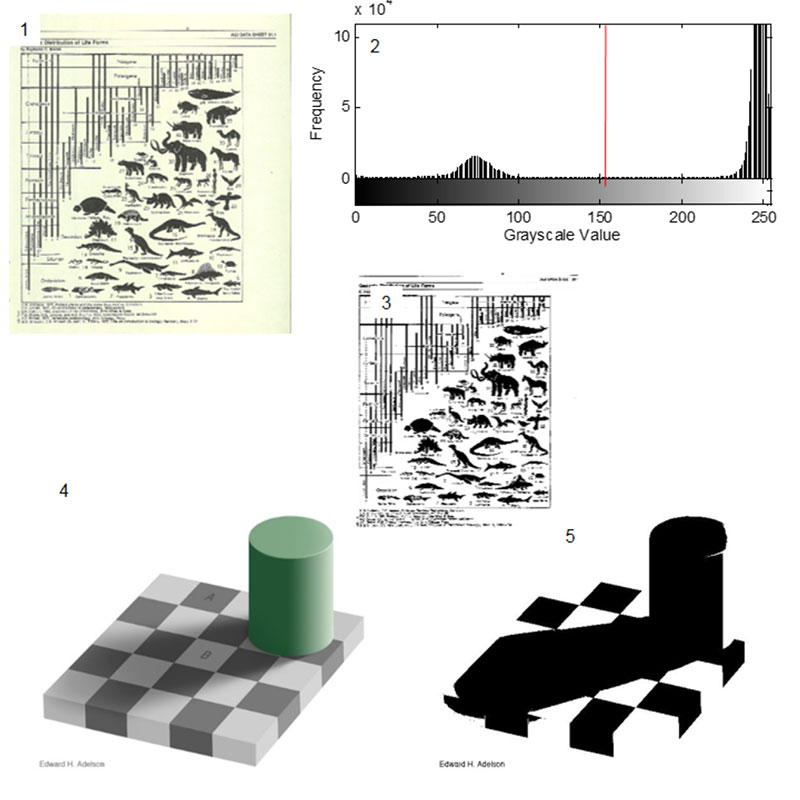
FIGURE 5. Applying local thresholding to the checker shadow illusion. The black checks and the white checks are correctly segmented.
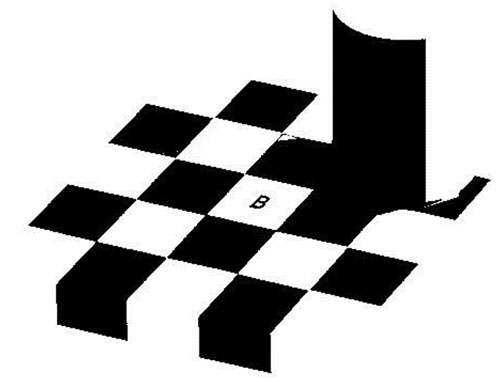

FIGURE 6. Result of the global thresholding of the image of a crinoid holdfast (see Figure 2.2). Without preprocessing there is a high level of noise (black spots).
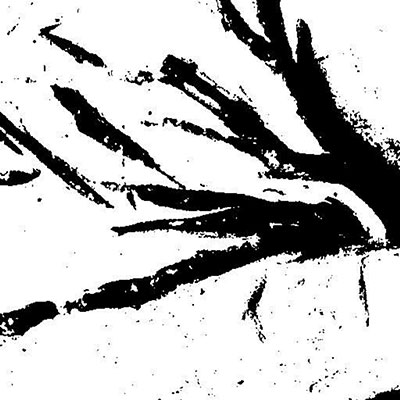

FIGURE 7. Effects of denoising. 1. The original grayscale image denoised. 2. How the denoising impacts the resulting thresholded image. Notice that there is less noise and irregular, porous edges than in Figure 6.
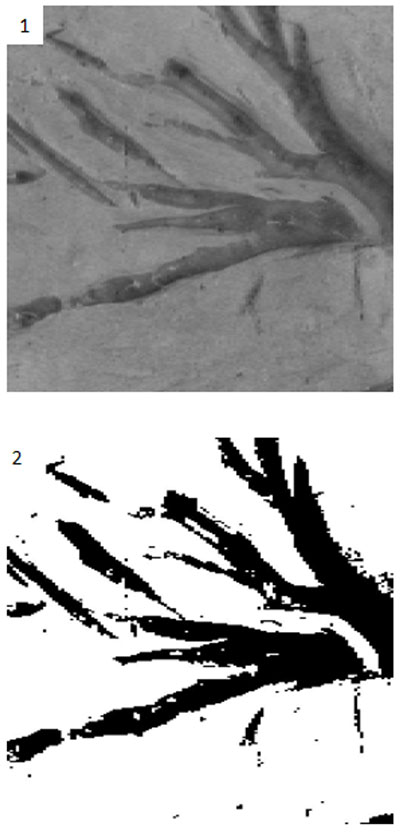
FIGURE 8. Connected component analysis. 1. The original binary image produced by preprocessing and segmentation of the tile shown in Figure 2.2. False color image of 8.1 showing the five largest objects in the image. Connected component analysis allows us to identify and remove the smaller objects. 3. Binary image after removing the smaller objects.
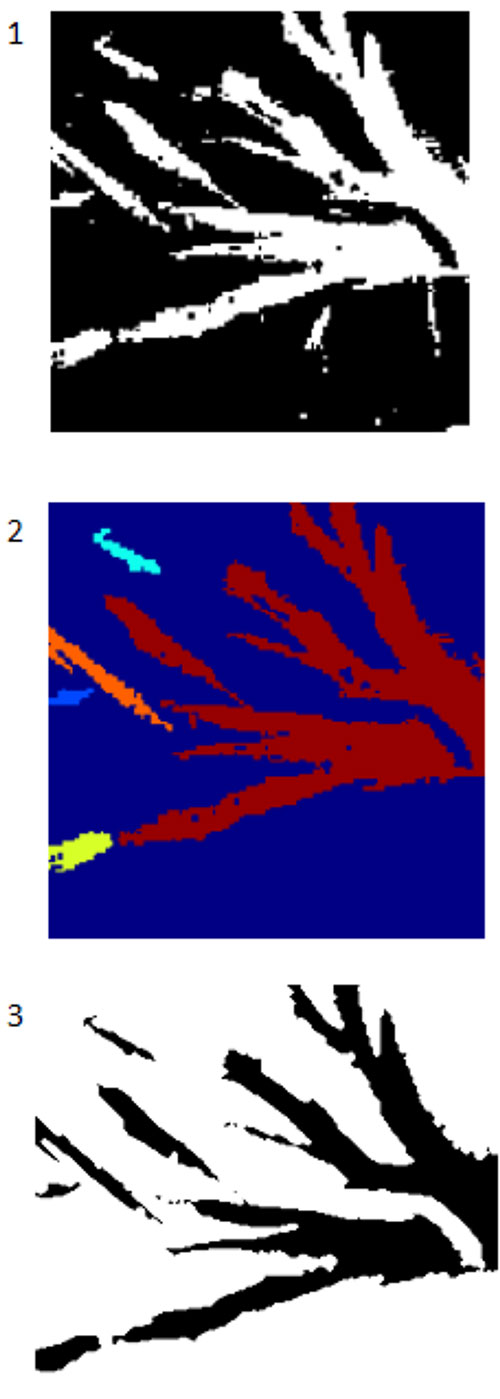

FIGURE 9. The effects of postprocessing using CC analysis and binary morphology. 1. CC analysis was applied to the denoised thresholded image in Figure 8. CC analysis has removed most of the remaining artifacts. 2. Once CC analysis is complete, binary image morphology can be used to smooth the edges of the image which better aligns with the real fossil image. 3. By applying another form of binary image morphology, an outline of the perimeter of the segmented image can be produced and the quality of the segmentation compared to the real specimen. Here we can see qualitatively that the match is very high.
FIGURE 10. Definitions of morphometric parameters. Each branch is defined as a link between nodes (circles). Each node has an order determined by the number of nodes from the stem nodes (purple). First order nodes (red) have no nodes separating them from the stem. Second order nodes (blue) have first order nodes separating them from the stem. Third order nodes (yellow) have second order nodes separating them from the stem. The node angle is the branching angle as defined by the next order nodes. The area is the area of the branch between the nodes, which are bounded by lines perpendicular to the bisector of the node angle.
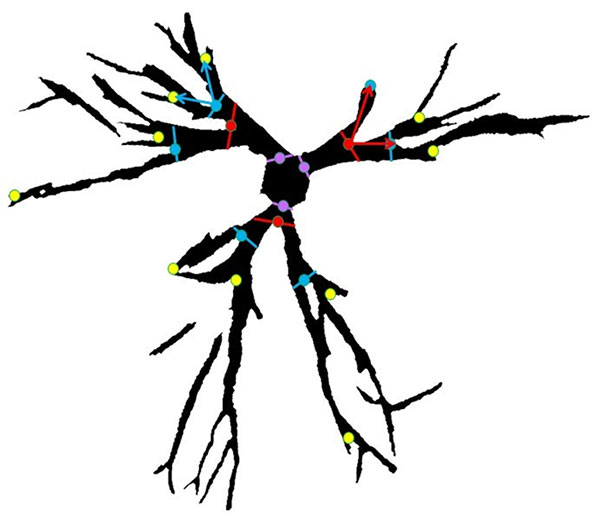
FIGURE 11. 1. Original scanned image of the bivalve Modiola from plate 8 of Murchison (1839) obtained via Google Books (TM). The digitized copy shows notable foxing. 2. Image reconstructed with thresholding of the green portion of the RGB image. This fast method of cleaning up the image for presentation and publication has removed the stains from the image.
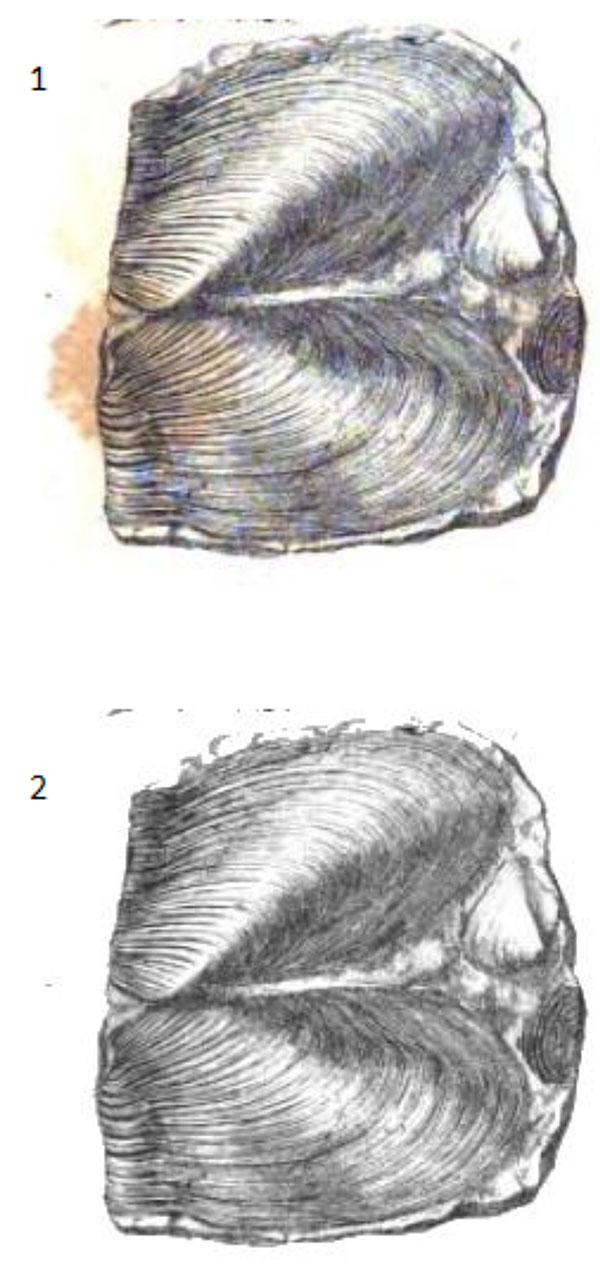
FIGURE 12. The effects of the techniques described in the paper on a graptolite rhabdosome from the University of Illinois at Chicago teaching collection, Silurian, Wales. Scale bar equals 5 mm. Processing was done on the original JPEG image. 1. Original image. 2. Segmented image.
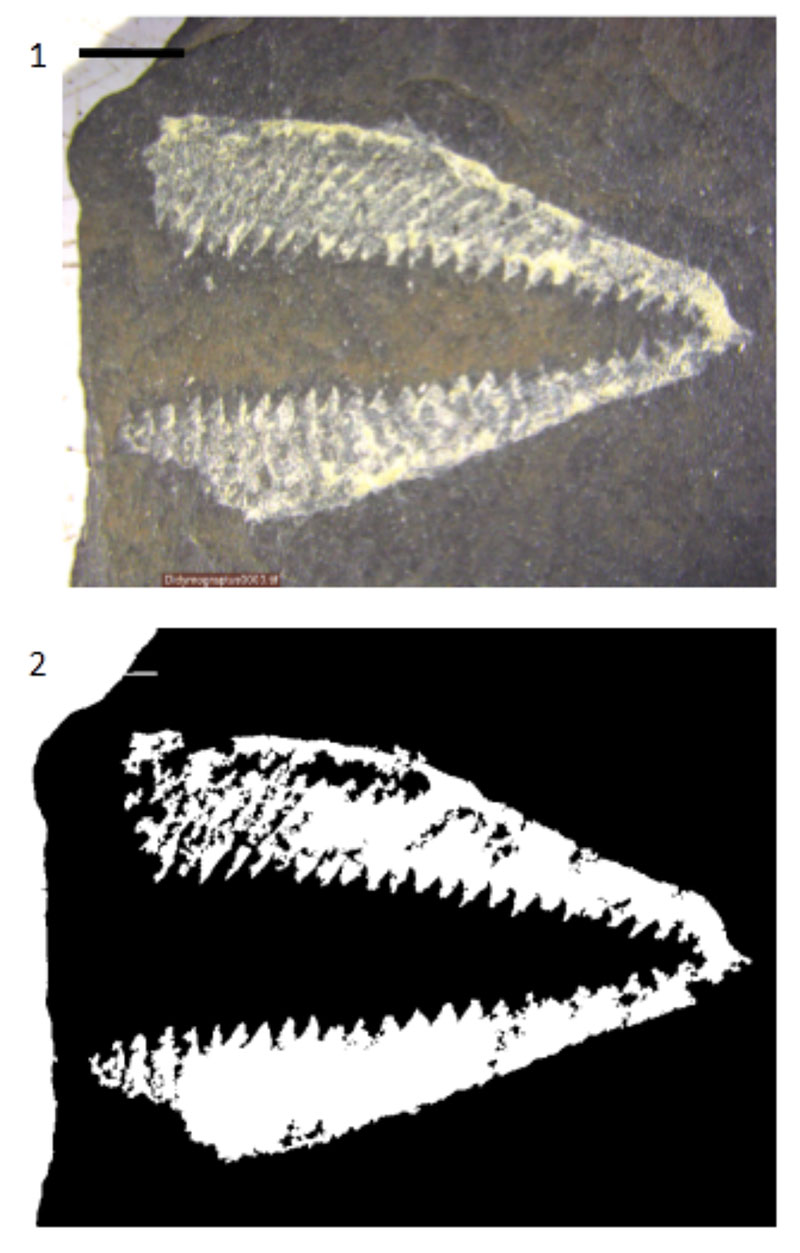

TABLE 1. Results of denoising a single branch and entire crinoid in a raw image. Wavelet shrinkage refers to the minimum nonzero level coefficients in the wavelet decomposition. An M file containing the method of wavelet shrinkage used here as well as calculating accuracy and MCC are in the supplementary materials..
TABLE 2. Morphometrics of crinoids. The length, distance to center (dist to center), width, and area are for each branch. The angle is the angle of the branch originating from each node.
| 86 | 53 | 74 | 53 | 102 | 76 | 74 | 56 | |
| 0.81 | 0.91 | 1.92 | 0.99 | 0.72 | 0.71 | 1.37 | 1.18 | |
| 1.26 | 2 | 2.39 | 2.9 | 1.33 | 1.73 | 1.56 | 2.73 | |
| 0.54 | 0.41 | 0.45 | 0.25 | 0.52 | 0.35 | 0.4 | 0.21 | |
| 0.45 | 0.32 | 0.63 | 0.19 | 0.4 | 0.17 | 0.36 | 0.24 | |
 Chris M.E. Honeycutt
Chris M.E. Honeycutt
Department of Laboratory Sciences
Jefferson Community College
1220 Coffeen Street
Watertown, New York 13601
USA
cebey@sunyjefferson.edu
Chris Ebey Honeycutt has been a faculty member at Jefferson Community College (SUNY Jefferson) since August 2013. The bulk of her publications have focused on applying mathematics and computer science to problems in paleontology, particularly ichnology and enigmatic fossils such as Paleodictyon. Her BS at the University of Illinois at Chicago was in mathematics, while her MS at the University of Illinois was in Earth and Environmental sciences. Her Ph.D. at the University of Southern Denmark was in biology, focusing on mathematical models of Proterozoic biogeochemistry, the Canfield Ocean, and carbon isotope fractionation.

Roy Plotnick  PE Emeritus Editorials Editor
PE Emeritus Editorials Editor
Department of Earth and Environmental Sciences
University of Illinois at Chicago
845 W. Taylor St.
Chicago, IL 60607
plotnick@uic.edu
Roy Plotnick is a professor in the Department of Earth and Environmental Sciences at the University of Illinois at Chicago, where he has been for more than thirty years since he received his doctorate at the University of Chicago. The inertia of his affiliation is in total contrast to the unpredictability of his scientific interests, which can be best be characterized as eclectic (some may some unfocused!) . He has published on eurypterids, arthropod taphonomy, functional morphology, the nature of wastebasket taxa, disparity, quantitative stratigraphy, trace fossils, and the applications of fractal and related methods in paleontology, stratigraphy, and landscape ecology. He is currently working on a remarkable Pennsylvanian paleokarst and cave fill, which preserves very early conifers. Roy has been associated with both PBDB and CHRONOS. Other interests include amateur astronomy, toy trains, and The Little Engine that Could.

 Fabien Kenig
Fabien Kenig
Department of Earth and Environmental Sciences
University of Illinois at Chicago
Science and Engineering South Building
(MC 186)
845 West Taylor Street
Chicago, Illinois 60607-7059
USA
fkenig@uic.edu
Fabien Kenig is an Earth scientist. He uses organic geochemistry and stable isotope analysis to answer questions in paleooceanography. A large part of his work focuses on biomarkers that can be used as climate proxies. He is currently a professor at the University of Illinois at Chicago in the department of Earth and environmental sciences.
Breaking free from the matrix: Segmentation of fossil images
Plain Language Abstract
Computers recognizing fossils: The ability of computers to recognize objects in images has evolved tremendously in the last few decades. The ability of computers to recognize objects is called computer vision. Computer vision can help paleontologists analyze images of fossils much faster. Our paper focuses on one type of computer vision technique, image segmentation. Here we use image segmentation to aid in identifying, isolating and measuring fossil crinoids. Crinoids are sea creatures with a 500 million year history. This research is part of a plan to build systems that can assist paleontologists in cataloging large specimen collections and identify fossils in remote locations.
Resumen en Español
Librarse de la matriz: la segmentación de imágenes de fósiles
El método tradicional para la separación de un fósil de su matriz es eliminar físicamente la matriz, arriesgándose a dañar el fósil. Las tecnologías no destructivas, como la tomografía computarizada (TC), permiten eliminar digitalmente la matriz. Sin embargo, estas técnicas tienen varias desventajas: se requiere mucho tiempo para realizar el proceso, precisan de preparadores o técnicos altamente cualificados, o requieren equipos costosos. La segmentación de imágenes, junto con la eliminación de ruido wavelet, proporciona una forma rápida, barata y no destructiva para separar digitalmente la matriz de los fósiles en imágenes estándar de cámaras digitales. Este método utiliza programas estándar y produce resultados que a continuación se pueden introducir en programas de análisis morfométrico o bien ser usados con las técnicas más tradicionales de segmentación para acelerar el proceso. Una comparación de métodos alternativos para la segmentación de imágenes y eliminación de ruido wavelet indica que el método de Otsu y otras técnicas de umbralización global funcionan mejor para este propósito. Este enfoque se ilustra aquí con imágenes de discos de fijación de Eucalyptocrinites. Las imágenes resultantes se utilizan en morfometría. Estos métodos tienen un gran potencial para los estudios que aprovechan el rápido crecimiento de imágenes digitales de museos disponibles en Internet.
Palabras clave: Imagen; segmentación; ondículas; extracción; crinoide; discos de fijación
Traducción: Enrique Peñalver
Résumé en Français
Se libérer de la matrice: segmentation d'images de fossiles
La méthode traditionnelle pour séparer un fossile de sa matrice est de supprimer physiquement la matrice, en risquant d'endommager le fossile. Des technologies non destructives, comme la tomodensitométrie (TDM), suppriment numériquement la matrice. Cependant, ces techniques ont plusieurs inconvénients: elles prennent du temps, impliquent des préparateurs ou des techniciens hautement qualifiés, ou nécessitent des équipements coûteux. La segmentation d'image, couplé avec du débruitage d'ondelettes, fournit un moyen rapide, peu coûteux et non destructif afin de séparer la matrice du fossile numériquement dans des images de caméras numériques standard. Cette méthode utilise des logiciels sur l'étagère et produit des résultats qui peuvent ensuite être entrée dans des logiciels d'analyse morphométrique ou utilisé pour accélérer les techniques de segmentation plus traditionnels. Une comparaison des méthodes alternatives pour la segmentation d'images et le débruitage d'ondelettes indiquent que la méthode d'Otsu et les autres techniques de seuillage mondiale fonctionnent mieux pour cette intention. Cette approche est illustrée ici avec des images de crampons d'Eucalyptocrinites. Les images obtenues sont utilisées pour la morphométrie. Ces méthodes ont un grand potentiel pour les études qui utilisent les catalogues en ligne d'images numériques des musées, une ressource en croissance rapide.
Mots-clés: image; segmentation; ondelettes; extraction; crinoïde; crampon
Translator: Kenny J. Travouillon
Deutsche Zusammenfassung
Befreiung aus der Matrix: Segmentierung von fossilen Bildern
Die traditionelle Methode ein Fossil aus der Matrix zu separieren, ist diese physikalische zu entfernen, wobei jedoch Beschädigungen riskiert werden. Nicht-destruktive Methoden wie Computertomografie (CT) entfernen Matrix digital. Diese Techniken haben jedoch vielerlei Nachteile: sie sind zeitaufwändig, erfordern hoch qualifizierte Präparatoren oder Techniker oder benötigen teure Ausrüstung. Bildsegmentation verbunden mit Wavelet-Transformation bietet einen schnellen, kostengünstigen und zerstörungsfreien Weg, um Matrix mit Bildern üblicher Digitalkameras digital vom Fossil zu separieren. Diese Methode benutzt serienmäßig produzierte Software und die Ergebnisse können dann in Software für morphometrische Analysen eingegeben werden oder dafür genutzt werden traditionellere Segmentationstechniken zu beschleunigen. Ein Vergleich von alternativen Methoden zur Bildsegmentierung und der Wavelet-Transformation zeigt, dass Otsu's Methode und andere globale Schwellentechniken am besten zu diesem Zweck geeignet sind. Dieser Ansatz ist hier mit Haftwurzeln von Eucalyptocrinites illustriert. Die resultierenden Bilder wurden für morphometrische Untersuchungen genutzt. In diesen Methoden steckt großes Potential für Untersuchungen welche die rasch anwachsende Ressource der digitalen Online-Bilderkataloge der Museen nutzen.
Schlüsselwörter: Bild; Segmentierung; Wavelets; Extraktion; Crinoide; Haftwurzel
Translator: Eva Gebauer
Arabic

Translator: Ashraf M.T. Elewa
-
-
-
Review: The Princeton Field Guide to Mesozoic Sea Reptiles
 The Princeton Field Guide to Mesozoic Sea Reptiles
The Princeton Field Guide to Mesozoic Sea ReptilesArticle number: 26.1.1R
April 2023


 Poster Winners 2024
Poster Winners 2024