 Computer simulation of the evolution of foraging strategies: application to the ichnological record
Computer simulation of the evolution of foraging strategies: application to the ichnological record
Article number: 1.2.5A
Copyright Palaeontological Association, 1 August 1998
https://doi.org/10.26879/98005
Plain-language and multi-lingual abstracts
PDF version
Submission: 7 April, 1997 Acceptance: 20 May, 1998
ABSTRACT
The ichnological record can provide invaluable insight into the evolution of behaviour. Much of the current work in so-called "artificial life" and artificial neural networks is applicable to ethological paleobiology. Some preliminary experiments in this direction are presented here. A computer generated community of detritus feeders was simulated with the individuals in each generation being subjected to selection based on the success of their feeding strategies. Mutation and sexual reproduction are also simulated, resulting in an evolutionary process that produces increasingly more advanced simulated neural control systems. The resulting "virtual trace fossils" can be compared with well-known, naturally occurring trace fossils and the fossil record of behavioural evolution. Though this study is not intended as an exact replication of natural processes, such simulations may aid theory formation, and can be a useful educational tool.
Oyvind Hammer, Paleontological Museum, Sars gt.1, N-0562 Oslo, Norway oyvindha@notam.uio.no
Keywords: Ichnology, evolution, simulation
Final citation: Hammer, Oyvind. 1998. Computer simulation of the evolution of foraging strategies: application to the ichnological record, Palaeontologia Electronica Vol. 1, Issue 2; 5A; 21p. https://doi.org/10.26879/98005
palaeo-electronica.org/content/1998-2/627-computer-simulation
INTRODUCTION
Trace fossils provide one of the few windows paleobiologists have on the evolution of animal behaviour. Feeding trails of presumed metazoan origin are known from the Vendian or even earlier (Fedonkin 1994), and the marine ichnological record of the Phanerozoic shows long-ranging trends of increasing efficiency of horizontal deposit feeding (Seilacher 1977). In this paper, some preliminary attempts at simulating the evolution of such behavioural patterns are described.
The role of computer simulation in ichnology, as in many biological subjects, can easily be misunderstood. Any similarity between the morphology of natural and simulated traces must be regarded as somewhat coincidental. One must take care not to overinterpret or otherwise assign too much weight to such occurrences. In this study the intention is not to simulate every aspect of the trace maker and its environment, which is, of course, impossible. Quantitative testing of behavioural evolutionary hypotheses using computer experiments is therefore not a valid approach. Still, computer simulation of the evolution of behavioural patterns can be a useful exercise because of the general insights it may provide. The experiments described below indicate, for example, that complex behaviour can arise from combinations of simple behavioural programs. Accordingly, any inference of correlation between the morphological complexity of a trace and the inferred complexity of the underlying behavioural program must be regarded with caution. In addition, if phenomena like adaptive radiations or punctuated equilibria arise in simulations with stable environments and without geographical isolation, this may give rise to questions of general significance without providing much evidence for or against the operation of particular natural processes. Finally, such simulations may have educational value through the comparison of simulated results with the actual fossil record.
ARTIFICIAL LIFE
The methods of Artificial Life (AL) are now well known to computer scientists (see Langton 1995 for an overview), but they have rarely been applied to paleontology. The key idea in current AL research is to simulate some aspects of natural evolution in the computer. This is done for two main reasons: (1) AL experiments can give insight into natural processes, and (2) AL techniques can be used for practical purposes in engineering.
Besides setting up the basic mechanisms of evolution (mutation, reproduction and selection), the most important element in the design of an AL system is the specification of the fitness criteria; that is, the method for determining the viability of a newly grown individual. Some examples of such criteria are: 1) the ability of simulated ants to find food, avoid obstacles and cooperate; 2) the ability of competing individuals to fight for resources and to eat, but not be eaten. "Ecosystems" with several trophic levels can emerge in such experiments (Yaeger 1994); 3) the ability of a computer virus to penetrate computer security systems; and 4) the ability of a computer program to solve a given problem. Note that an "individual" is not necessarily a simulated biological entity, but can just as well be a simulated machine or computer program with properties that are to be optimized. "Artificial Life" (AL) is becoming one of the most dynamic and promising fields of computer science, partially displacing the position previously held by "Artificial Intelligence" (AI).
In a very early work in the field of AL, F. Papentin simulated a population of "worms", producing feeding patterns that could be compared with natural trace fossils (Papentin 1973, Papentin and Roder 1975, see also Raup and Seilacher 1969). Papentin modelled the organisms using six simple ethological rules, and limited them to turning in 90 degree angles. However, these parameters could mutate and evolve, thus changing the rules. With this approach, efficient foraging strategies like tight meandering and spiraling emerged. The current study is a continuation of these experiments, but no presuppositions about behaviour are made. Rather, the organisms are allowed to freely develop an augmented artificial neural network as their control structure. Similar "foraging" simulations are in fact important model problems in modern AL research (Miglino et al. 1996, Langton 1995), but comparison with the trace fossil record has not been an important element. The "food" is generally distributed in discrete, widely-spaced packages, which does not encourage the evolution of natural-looking trace fossils.
EXPERIMENTAL SETUP
A population of simulated detritus feeders is living in a two-dimensional simulated world consisting of both barren areas and areas rich in nutrients. The population size is limited to 400 individuals, to avoid excessive computation time. The arena is a quadratic grid of 50 by 50 unit squares, giving both acceptable resolution and computational efficiency. Three elliptical "food patches" are placed with their centers at pseudorandom positions (uniform distribution). The ellipses are generated by stretching circles with unit radius by pseudorandom factors in the horizontal and vertical directions separately, giving major and minor radii between 8 and 15 units. The food patches are given new positions and scalings between each generation, to discourage the organisms from specializing to a specific geography.
Moving at a constant speed of one unit distance per time step, the "creatures" pick up any food present and leave behind a "mucus trail." All individuals are given a constant lifespan of 400 time steps, and are selected for their feeding success during this interval. Ideally, they should then find efficient ways to steer, avoiding barren areas and their own trail as much as possible. In each generation, the 80 individuals with highest fitness values (number of food units eaten) are automatically selected for reproduction. Among these, reproductive success is further controlled statistically from their fitness, by setting the probability of selecting a parent to its normalized fitness value. Reproduction is controlled by the program to keep the population size constant. Technically, this is done by keeping a "slot" for each of the possible 400 organisms. When an organism is removed because of low fitness value, its slot becomes vacant. In the next generation, all vacant slots are filled with new organisms. Reproduction involves both pseudorandom mutation and exchange of genetic material between the two parents (see below).
The genotype and phenotype are undistinguishable in this system, as they consist of the same data structure. A network of "modules" are connected together to form a control system in the form of a directed graph, possibly with feedback loops. This network can be compared with a normal artificial neural network, but includes additional module types that have little biological justification in the hope that this will lead to complex strategies after few generations. A total of 25 rather arbitrarily chosen module types are available, including sensors, summers, multipliers, nonlinear transfer functions, memory, oscillators, etc. (Table 1). Each input has an associated weight factor. A single output controls the magnitude and direction of turning, much like the driving wheel in a car. The module types were mostly taken from Sims 1994, who used a similar method to successfully evolve control structures for "swimming" and "jumping" in simulated organisms.
 Mutation involves pseudorandom changes in the input weightings in the modules, addition and removal of modules, and addition and removal of communication links ("axons") between modules.
Mutation involves pseudorandom changes in the input weightings in the modules, addition and removal of modules, and addition and removal of communication links ("axons") between modules.
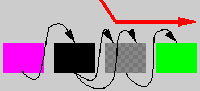
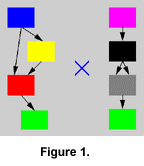
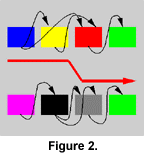
Parts of the network can be interchanged between two individuals during mating in a process that can be compared with chromosomal crossover. This is accomplished by arranging the modules of the parents in two linear rows, and combining segments of the rows into a new row representing the child (Figure 1, Figure 2, and Figure 3). The segments are chosen by copying modules from one row and switching to the other row with a probability of 0.12 (value arbitrarily chosen) for each traversed module. If the parents are dissimilar, such crossover will normally produce a highly dysfunctional control structure, because modules are integrated into a functional whole and are not likely to function in another context. Such an effect might conceivably lead to the formation of "species," but this has not been investigated in the current experiments.
AN EXAMPLE CONTROL NETWORK
 The evolved control networks are accessible for study, but deciphering how they actually work is more difficult. Compared with computer programs written by human programmers, evolved algorithms are less likely to refer to the resulting behaviour in a modular, linear and one-to-one way.
The evolved control networks are accessible for study, but deciphering how they actually work is more difficult. Compared with computer programs written by human programmers, evolved algorithms are less likely to refer to the resulting behaviour in a modular, linear and one-to-one way. 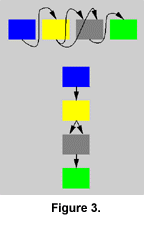 What we believe we observe as one distinct ethological "rule," may in fact be an emergent product of a large nonlinear network with feedbacks and memory. Conversely, a small subnetwork may be responsible for several different behaviours in different contexts. Consequently, the correlation between complexities of the behavioural program and the trace it produces is not necessarily strong.
What we believe we observe as one distinct ethological "rule," may in fact be an emergent product of a large nonlinear network with feedbacks and memory. Conversely, a small subnetwork may be responsible for several different behaviours in different contexts. Consequently, the correlation between complexities of the behavioural program and the trace it produces is not necessarily strong.
As an example, consider a network that was observed in one simulation run (Figure 4). One of the modules in the inventory is a food sensor that gives information about the presence of food in a given direction ("FSENSOR" in Table 1). The creatures were expected to acquire a set of such sensors, set up to sense the state of the environment in different directions (e.g., to the left, to the front and to the right). 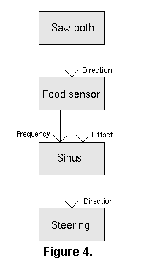 However, there is another module called a "sawtooth oscillator," that gives value increasing linearly to a given level and then decreasing to zero again ("SAW" in Table 1). This creature had evolved a network where such an oscillator was coupled to the direction parameter of a single sensor, creating a sweeping sensor that could report the environmental state in all directions successively, much like a radar. This may be an obvious idea in hindsight, but nevertheless a clever solution that the programmer did not foresee. Further, this organism walked straight forward until a food patch was encountered, and then turned to a meandering behaviour. When the outer limit of the patch was reached, the organism switched to walking with constant curvature, which might bring it back into the patch. If it did not return to the patch, the curvature was exponentially decreased towards straight walking again. This searching phase could bring the organism in contact with another patch where the meandering resumed. All this must be regarded as rather elaborate behaviour given the small size of the control network.
However, there is another module called a "sawtooth oscillator," that gives value increasing linearly to a given level and then decreasing to zero again ("SAW" in Table 1). This creature had evolved a network where such an oscillator was coupled to the direction parameter of a single sensor, creating a sweeping sensor that could report the environmental state in all directions successively, much like a radar. This may be an obvious idea in hindsight, but nevertheless a clever solution that the programmer did not foresee. Further, this organism walked straight forward until a food patch was encountered, and then turned to a meandering behaviour. When the outer limit of the patch was reached, the organism switched to walking with constant curvature, which might bring it back into the patch. If it did not return to the patch, the curvature was exponentially decreased towards straight walking again. This searching phase could bring the organism in contact with another patch where the meandering resumed. All this must be regarded as rather elaborate behaviour given the small size of the control network.
RESULTS
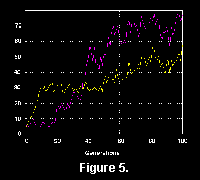 The simulator, as perhaps evolution itself, can be seen as an iterative optimizing system for a nonlinear function (fitness). As is well known from such systems (e.g., Miglino et al. 1996), the program can get stuck on a local maximum for a long time before a new "invention" breaks through the selection process (Figure 5, Figure 6 , Figure 7). Therefore, there are long periods of little activity which we can refer to as stasis, though of course without implying that the phenomenon has exactly the same reasons as in nature.
The simulator, as perhaps evolution itself, can be seen as an iterative optimizing system for a nonlinear function (fitness). As is well known from such systems (e.g., Miglino et al. 1996), the program can get stuck on a local maximum for a long time before a new "invention" breaks through the selection process (Figure 5, Figure 6 , Figure 7). Therefore, there are long periods of little activity which we can refer to as stasis, though of course without implying that the phenomenon has exactly the same reasons as in nature.
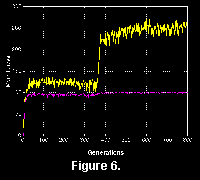 Most often, a flurry of activity occurred in the first few generations, causing an "adaptive radiation" with many extravagant experiments. Soon however, most of the forms were decimated as the few most efficient grazers filled the living space. One of the most dramatic events in these simulations was when the first creature discovered an advantageous use of sensors, representing a key innovation. The "genes" of this individual quickly spread and disrupted the existing blind (but often well diversified) communities. In the run shown in Figure 6 it can be seen that a significant subpopulation had acquired a food sensor some time before it was actually connected in a highly useful way. The inclusion of a sensor element was therefore probably initially a character with limited adaptive value, but represented a "pre-adaptation" for an important breakthrough.
Most often, a flurry of activity occurred in the first few generations, causing an "adaptive radiation" with many extravagant experiments. Soon however, most of the forms were decimated as the few most efficient grazers filled the living space. One of the most dramatic events in these simulations was when the first creature discovered an advantageous use of sensors, representing a key innovation. The "genes" of this individual quickly spread and disrupted the existing blind (but often well diversified) communities. In the run shown in Figure 6 it can be seen that a significant subpopulation had acquired a food sensor some time before it was actually connected in a highly useful way. The inclusion of a sensor element was therefore probably initially a character with limited adaptive value, but represented a "pre-adaptation" for an important breakthrough.
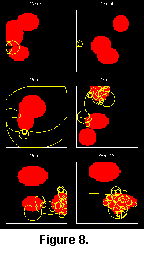
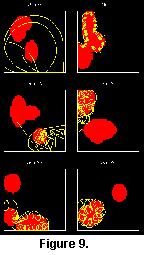
 Some of the patterns observed in the fossil record could be seen in the simulations, in particular the general trend towards more efficient grazing patterns (Seilacher 1977). From unsystematic scribbling, the creatures evolved towards meandering, spiralling and edge following. However, some natural patterns have not arisen in the simulations. In particular, tight, labyrinthic meanders (Helminthoides) never evolved. This may be due to the way the environment was simulated, or perhaps the simulations were not allowed to run for a sufficient number of generations (Figure 8 and Figure 9).
Some of the patterns observed in the fossil record could be seen in the simulations, in particular the general trend towards more efficient grazing patterns (Seilacher 1977). From unsystematic scribbling, the creatures evolved towards meandering, spiralling and edge following. However, some natural patterns have not arisen in the simulations. In particular, tight, labyrinthic meanders (Helminthoides) never evolved. This may be due to the way the environment was simulated, or perhaps the simulations were not allowed to run for a sufficient number of generations (Figure 8 and Figure 9).
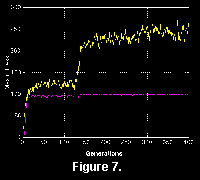
Formal statistics and comparison of simulation results are not given here, for several reasons. Since a single run can take an hour or more of computing time large numbers of simulations are not presently practical. Secondly, different runs with slightly different parameters can give very different results (compare Figure 6 and Figure 7), perhaps showing the importance of "historical contingency" (Gould 1989) in these simulations. The large variation in results means that comparing different parameters would necessitate very large numbers of simulations to provide meaningful statistics. Thirdly, meaningful formal measures on the morphology of traces are not known to the author. Finally, and most importantly, there is a danger of over-interpreting these extremely simplified simulations.
USING STANDARD NEURAL NETWORKS
 A valid objection to the experiments presented here is that many of the module types are rather artificial from a biological point of view. Sinus oscillators and square root functions hardly exist as discrete entities in real neural systems. Considering the success in recent years of mimicking real neural networks in computer simulations (Haykin 1994), simulations were done where the inventory of modules was limited to sensors and a simulated neuron. Such a neuron consisted of a number of inputs (maximally three) that were weighted and summed together, and a nonlinear threshold transfer function:
A valid objection to the experiments presented here is that many of the module types are rather artificial from a biological point of view. Sinus oscillators and square root functions hardly exist as discrete entities in real neural systems. Considering the success in recent years of mimicking real neural networks in computer simulations (Haykin 1994), simulations were done where the inventory of modules was limited to sensors and a simulated neuron. Such a neuron consisted of a number of inputs (maximally three) that were weighted and summed together, and a nonlinear threshold transfer function:
 The weights wi and the transfer function parameter k were subject to mutation, but were fixed during the lifespan of a single organism. No learning could therefore occur as all values were inherited. The topology of the network was freely evolvable as before, and networks with feedback loops would allow memory mechanisms and oscillators to develop.
The weights wi and the transfer function parameter k were subject to mutation, but were fixed during the lifespan of a single organism. No learning could therefore occur as all values were inherited. The topology of the network was freely evolvable as before, and networks with feedback loops would allow memory mechanisms and oscillators to develop.
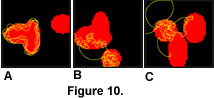
The exclusion of non-biological modules did not stop the evolution of some complex and efficient strategies, as shown in Figure 10.
A complete program for UNIX computers (only tested on SGI) is given here. Listings of the particular programs with their many parameters that were used to produce the examples in this paper are available via FTP at ftp://www.notam.uio.no/pub/SNIL/.
FUTURE WORK
 Many experiments can be carried out in AL systems like this. For example, what happens if the environment is suddenly changed? How do the "genes" spread? Geography can also be introduced, allowing experiments in migration, allopatric speciation, etc. Three-dimensional burrowing can be simulated to see if traces like Zoophycos and Chondrites will develop. The evolution of trace fossils should be a useful model problem in evolutionary biology and AL, because of the simple fitness criteria, the relatively simple neural control systems of the organisms involved and the availability of a fossil record for comparison.
Many experiments can be carried out in AL systems like this. For example, what happens if the environment is suddenly changed? How do the "genes" spread? Geography can also be introduced, allowing experiments in migration, allopatric speciation, etc. Three-dimensional burrowing can be simulated to see if traces like Zoophycos and Chondrites will develop. The evolution of trace fossils should be a useful model problem in evolutionary biology and AL, because of the simple fitness criteria, the relatively simple neural control systems of the organisms involved and the availability of a fossil record for comparison.
ACKNOWLEDGMENTS
Thanks to Whitey Hagadorn for all his suggestions and ideas.
REFERENCES
Fedonkin, M. A. 1994. Vendian body fossils and trace fossils. In Bengtson, S. (ed.), Early life on earth, Nobel symposium no. 84. Columbia University Press.
Gould, S. J. 1989. Wonderful Life: The Burgess Shale and the Nature of History. W. W. Norton & Company.
Haykin, S. 1994. Neural networks. Macmillan College Publishing Company, New York.
Langton, C. G. (ed.). 1995.Artificial Life - An Overview. MIT Press.
Miglinio, O., Nolfi, S. and Parisi, D. 1996. Discontinuity in Evolution: How Different Levels of Organization Imply Preadaptation. In Belew, R. K. and Mitchell, M. (eds.), Adaptive Individuals in Evolving Populations: Models and Algorithms. Addison-Wesley.
Papentin, F. 1973. A Darwinian evolutionary system. Journal of Theoretical Biology, 39:397-415 and 431-445.
Papentin, F, and Roder, H. 1975. Feeding patterns: The evolution of a problem and a problem of evolution. Neues Jahrbuch fur Geologie und Palaontologie, Monatshefte 1975:184-191.
Raup, D. M. and Seilacher, A. 1969. Fossil foraging behaviour: Computer Simulation. Science, 166:994-995.
Seilacher, A. 1977. Evolution of Trace Fossil Communities, p. 359-376. In Hallam, A. (ed.), Patterns of evolution, as illustrated by the fossil record. Elsevier.
Sims, K. 1994. Evolving Virtual Creatures. Computer Graphics, Annual Conference Series, (SIGGRAPH '94 Proceedings), July 1994:15-22.
Yaeger, L. 1994. Computational Genetics, Physiology, Metabolism, Neural Systems, Learning, Vision, and Behavior or PolyWorld: Life in a New Context. In Langton, C. et al. (eds.), Artificial Life III. Addison-Wesley.

